Your Genome Was Never in the Study, Only 1 in 600: Why Arab DNA Is the Biggest Blind Spot in Modern Medicine
March 15, 2026

Here's a number that should bother you: 1 in 600.
That's how many participants in published genome-wide association studies (GWAS) - the large-scale research that powers modern genetic medicine - are of Arab ancestry. One in six hundred. Meanwhile, Arabs make up nearly 5-6% of the world's population, roughly 500 million people. [1]
To put that another way: 523 out of 600 participants in those same studies are European. Arab DNA is almost missing from the genomic data that underpins modern risk scores, drug targets, and “personalized” medicine.
This isn't a minor statistical footnote. It's a fault line running through the foundation of precision medicine, and for anyone seeking a genetic diagnosis in the Arab world, it has very real consequences.
The Map Has a Blank Spot
Think of global genomic databases like a map. The more genomic data you feed in from a population, the more detailed and accurate that map becomes for people from that background. Over decades, researchers have drawn an extraordinarily detailed map of European genetics, followed by East Asian genetics, then South Asian genetics and African Genetics.
For Arab genetics? Much of the map is still blank.
The GWAS Catalog, which is the world's largest repository of genetic association studies, contains zero genomes from Algeria, Iraq, Libya, Oman, Somalia, and Syria. Zero. [1] And the imputation panels that clinicians worldwide use to fill in missing genetic data perform up to 61% worse when applied to Arab individuals compared to a population-specific Arab reference. [2]
That 61% isn't a rounding error. It's the difference between a confident clinical answer and a frustrating "we're not sure."
Arab Genetics Isn't Just "Missing Data", It's Genuinely Distinct
Here's where it gets scientifically interesting.
Arab genetic architecture isn't simply European genetics with a different label. It is its own genomic landscape, shaped by thousands of years of a unique demographic history, geographic isolation, and cultural practices that have no real parallel in the populations that dominate global genomic databases.
A few things set it apart:
Deep founder lineages and unique Arab variants. Analysis of over 6,000 Qatari whole genomes identified five major Arab ancestry clusters, each with their own unique variant profiles. Peninsular Arabs trace their lineage to the earliest Eurasian populations, carrying genetic signatures tied to splits that occurred 12-20 thousand years ago, variants that simply don't exist in standard European or East Asian reference panels.[2] The Qatar Genome Program dataset alone contained nearly double the number of variants compared to the Haplotype Reference Consortium, despite having 80% fewer samples. That's how different Arab genetic diversity really is.
Consanguinity and its genomic fingerprint. Marriage between relatives is significantly more common in Arab populations, estimated at 20-50% of unions across the Greater Middle East, compared to less than 0.2% in Western Europe and the Americas.[3] This isn't just a cultural fact. It leaves a measurable genomic signature: elevated runs of homozygosity, a higher burden of recessive genetic disease, and when studied properly reveal a uniquely powerful window into gene function. A 2026 study leveraging this feature identified over 180 knocked-out genes in the Qatari population, revealing essential genomic regions that outbred European cohorts could never have uncovered.[6]
The polygenic risk score problem. This one hits closest to the clinic. PRS tools, which are increasingly used to predict your risk of heart disease, diabetes, or cancer, are calibrated almost entirely on European data. When applied to Arabs, the minor allele frequency of causal variants is on average 7.6% lower than in European populations.[1] That gap systematically degrades predictive accuracy. And as PRS tools expand into clinical practice across the Arab world, a miscalibrated risk score isn't just imprecise, it's potentially harmful.
A 10-Year Diagnostic Odyssey
When the Reference Doesn’t Fit, Diagnosis Becomes a Decade-Long Journey Numbers are one thing. But behind every missed variant is a person waiting for an answer.
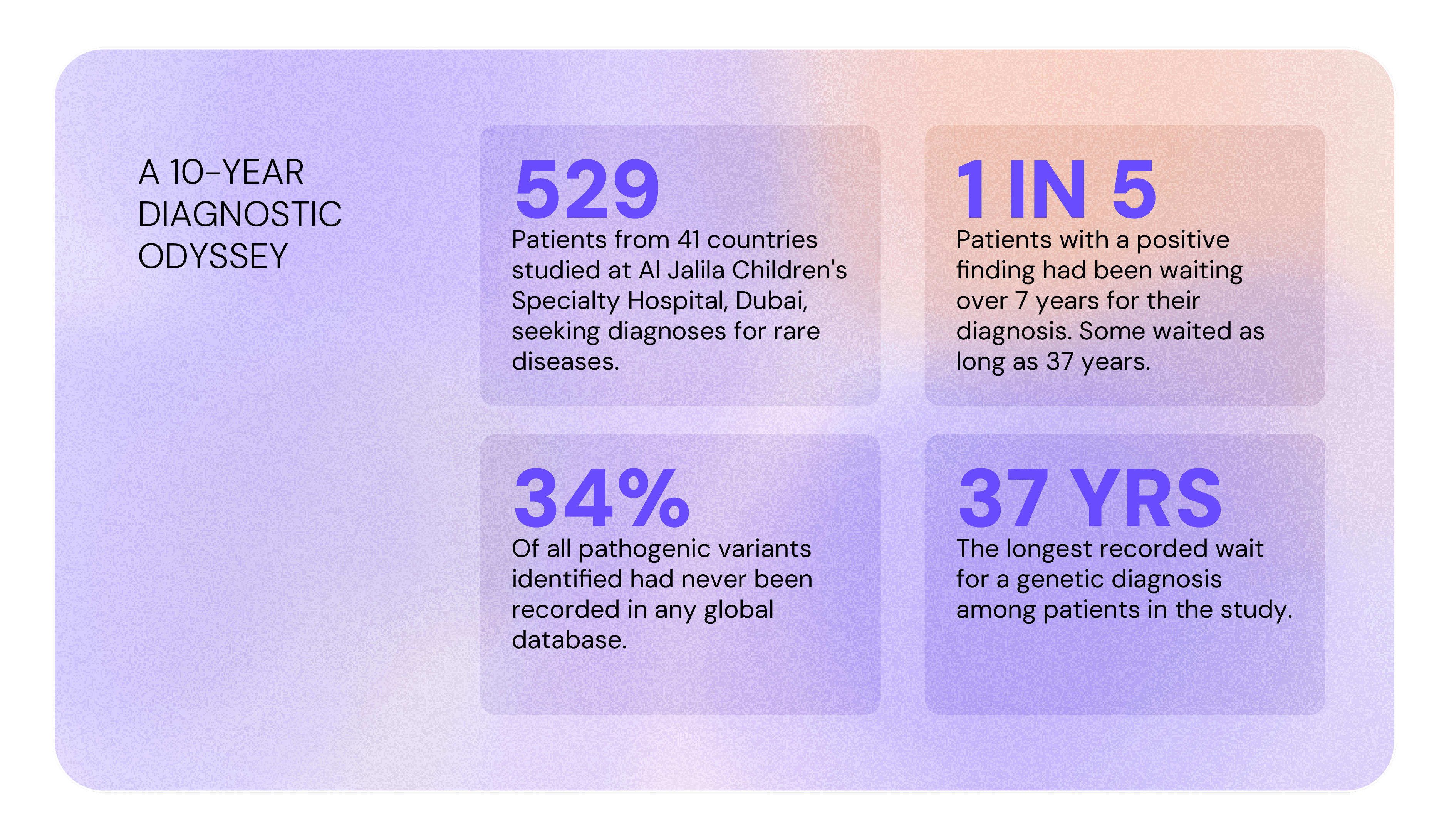

A study at Al Jalila Children's Specialty Hospital in Dubai followed 529 patients from 41 countries, mostly from the Arabian Peninsula, the Levant, and the broader MENA region, seeking genetic diagnoses for rare diseases. Nearly 1 in 5 patients with a positive finding had been waiting over 7 years for their diagnosis. Some waited as long as 37 years. Of all pathogenic variants identified, 34% had never been recorded in any global database.[4]
One patient, a man in his twenties, had spent over a decade cycling through hospitalizations with no working diagnosis. Whole exome sequencing finally identified a homozygous variant in the DOCK8 gene, diagnosing him with a life-threatening immunodeficiency. He was transferred to the NIH, found to have concurrent non-Hodgkin's lymphoma, and successfully treated with a stem cell transplant.[4]
Ten years. One variant. One database that finally contained enough Arab genomic data for his genome to be recognized.
Green Shoots
The tide is turning, slowly, but meaningfully.
The Qatar Genome Programme, launched in 2015, has sequenced over 45,000 whole genomes of Qatari citizens and residents, making it the largest contributor of Arab genomic data to global research to date, enabling the discovery of 24.6 million previously unknown variants and a population-specific imputation panel that substantially outperforms global tools for Arab rare variant classification.[2] The Emirati Genome Programme, launched in 2019 with the explicit goal of addressing Arab underrepresentation in global genomics, has collected over 815,000 samples as of 2025, one of the most ambitious national sequencing efforts anywhere in the world. [7] The 2025 Arab Pangenome then pulled it all into focus: over 111 million base pairs of Arab DNA entirely absent from the standard human reference genome, and 235,000 structural variants unique to Arab individuals that had never been catalogued anywhere.[5]
These are landmark achievements. But they come with an important caveat.
Qatar is one country. The Arab world is 22. The Levant, the Gulf, and North Africa each carry their own distinct genetic history, disease landscape, and reference gaps. A Jordanian patient, a Moroccan patient, an Iraqi patient: none of them are well-served by a reference built on a single Gulf population, however carefully constructed.[1] And across most of those countries, large-scale genomic research has barely begun.
The work is far from done.
The Patients Exist. What's Missing Is the Infrastructure.
Arab hospitals are already generating genomic data, including clinical exomes, sequencing runs, variant calls, every single day. The problem is not only a lack of data; It's also that the infrastructure to make it useful, locally, securely, against population-appropriate references, has largely not existed.
Closing the Arab genomic gap requires more than waiting for larger international studies to eventually include a MENA cohort as an afterthought. It requires building the capacity to generate, analyze, and interpret genomic data in-region. This must be done against databases that actually reflect Arab genetic diversity, in compliance with local data sovereignty laws, and with the clinical reporting pipelines to make findings actionable for the patients who need them most.
That's the work Bionl.ai is building toward. Alongside our bioinformatics platform, we're developing a MENA-focused patient cohort infrastructure: a privacy-first environment where regional hospitals can curate and contribute de-identified genomic datasets, and where researchers can finally access the population-specific data that has been missing from global science for decades. Not as a data broker. Not as another Western-centric marketplace with a MENA checkbox. As a platform built from the ground up for this region, its data, and its patients.
The science has been telling us for years what's needed. Some data is already there. What's arriving is the infrastructure to finally use it. Learn more →
References
[1] Bhattacharya R, et al. Massive underrepresentation of Arabs in genomic studies of common disease. Genome Medicine. 2023;15:99. https://doi.org/10.1186/s13073-023-01254-8
[2] Mohamad Razali R, et al. Thousands of Qatari genomes inform human migration history and improve imputation of Arab haplotypes. Nature Communications. 2021;12:5929. https://doi.org/10.1038/s41467-021-25287-y
[3] Scott EM, et al. Characterization of Greater Middle Eastern genetic variation for enhanced disease gene discovery. Nature Genetics. 2016;48:1071–1076.
[4] El Naofal M, et al. The Genomic Landscape of Rare Disorders in the Middle East. medRxiv. 2022. https://doi.org/10.1101/2022.09.17.22279590
[5] Ward N. The Missing 6 Percent: Why Arab DNA Is Changing the Future of Genomics. The Pathologist. October 2025.
[6] Falchi M, Fakhro K, et al. The biomedical landscape of genomic structural variation in the Qatari population. Nature Communications. 2026. https://doi.org/10.1038/s41467-025-67763-9
[7] Pearce N. More than 800,000 Emiratis contribute to UAE genome programme to boost health of the nation. The National. April 17, 2025. https://www.thenationalnews.com/news/uae/2025/04/17/more-than-800000-emiratis-contribute-to-uae-genome-programme-to-boost-health-of-the-nation/
Related Posts
- Genetics
- Research
The Effect of Genetics on Drug Metabolism
Genetics plays a key role in how individuals metabolize drugs, influencing both effectiveness and risk of side effects. Real-world cases like codeine, citalopram, and azathioprine highlight the clinical impact of tailoring treatments through genetics.
Aug 2023
Bayan Abu Alragheb
- Bioinformatics
- Research
Reproducibility in Bioinformatics Research with Bionl
Reproducibility, the cornerstone of scientific integrity, is the ability to duplicate the results of a scientific experiment or study. In the realm of bioinformatics research, reproducibility takes on a heightened significance due to the complexity of experiments, specialized software, and large datasets involved. This article explores the challenges and solutions to achieving reproducibility in bioinformatics research, with a special focus on how Bionl.ai can facilitate this crucial aspect.
Nov 2023